Astrophysical Data and Surveys
The Astronomical Data group develops and maintains advanced tools for the astrophysics community, especially ones for building probabilistic models and making precise measurements. It builds these tools by carrying out in-house data analysis projects that answer important scientific questions.
CCA
Astrophysical Disks and Jets
Astrophysical Gases and Fluids
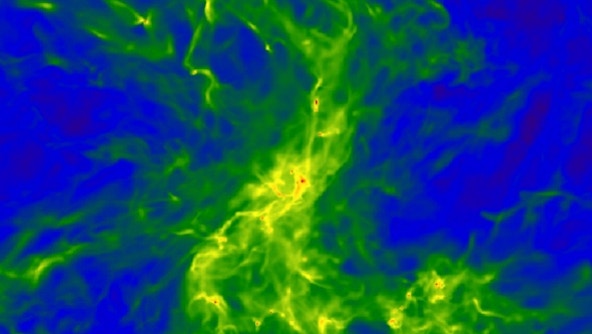
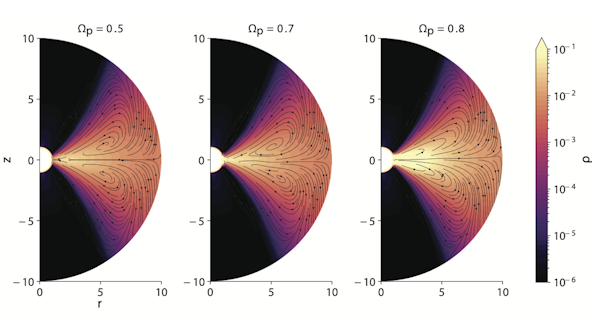
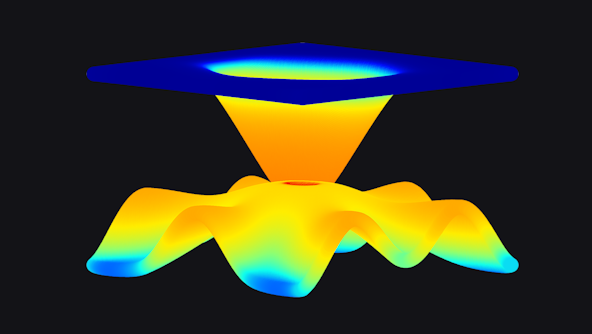
Astrophysical Gases and Fluids research at CCA is at the forefront of understanding the complexities surrounding galactic gas flows, stellar interiors, stellar atmospheres and magnetospheres, and accretion disks. Leveraging cutting-edge magnetohydrodynamic and kinetic particle-in-cell codes, CCA researchers delve deep into the multifaceted, multi-timescale and multi-wavelength signatures of astrophysical fluids.
CCA
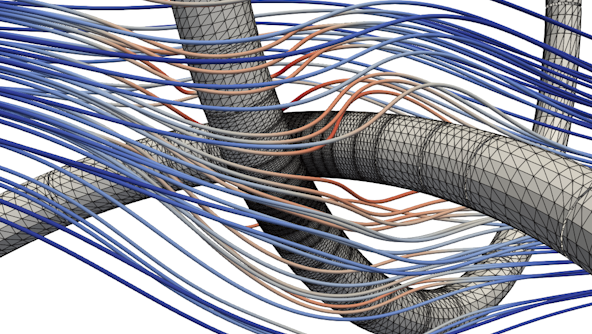
Biological Transport Networks
The Biological Flow Networks Group employs theoretical and computational methodologies to elucidate the underlying principles governing the evolution, development and functionality of biological flow networks, such as arterial, venous, and lymphatic systems and biological transport systems in general.
CCB
Biomolecular Design
The theory of protein folding explains how a disordered chain of amino acids spontaneously adopts a well-defined three-dimensional structure in water. The ultimate test of a theory is its successful application to the design of new physical systems with new, desired properties. In the Biomolecular Design Group, we apply the theory of protein folding to design new heteropolymers that fold into well-defined three-dimensional structures with new, useful functions, but which are built from exotic chemical building-blocks that go beyond the 20 canonical amino acids that make up natural proteins.
CCB
Biophysical Modeling
The Biophysical Modeling group focuses on the modeling and simulation of complex systems that arise in biology and soft condensed matter physics. Areas of interest include the dynamics of complex and active materials, as well as aspects of collective behavior and self-organization in both natural systems (e.g., inside the cell) and synthetic ones.
CCB
Compact Objects
CO
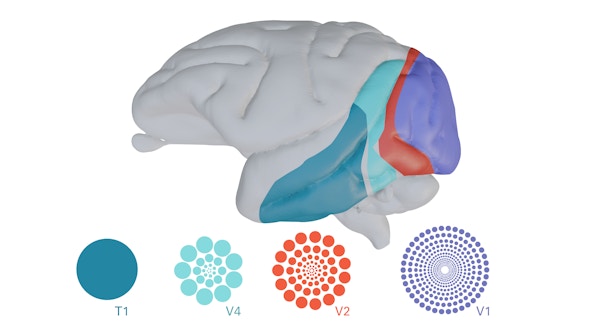
Computational Vision
We are interested in the analysis and representation of visual information, including empirical study of the structure of visual scenes, construction of mathematical theories for representation and processing that structure.
CCN
Cosmic Microwave Background
CDS
Cosmology
Researchers at the Center for Computational Astrophysics (CCA) stand at the forefront of advancing our understanding of the universe by melding cutting-edge machine learning and data science techniques with profound cosmological inquiries. In an era where cosmology is undergoing a transformative shift, this group spearheads the development of pioneering algorithms and innovative conceptual frameworks that are tailored to the increasingly complex cosmological datasets. Harnessing the collective power of ground-based and space-based telescopes furnished with progressively sensitive cameras and instruments, researchers at CCA are at the vanguard of endeavors such as SDSS, Vera Rubin Observatory, Euclid, SPHEREX, HIRAX and Roman Space Telescope.
CCA
Dark Matter in the Nearby Universe
Dealing with Dynamical Disequilibrium
Detection and Characterization
Developmental Dynamics
The Developmental Dynamics group combines experiments, theory and computing to elucidate the contributions of encoded genomic instructions and self-organizing physical mechanisms to embryonic development.
CCB
Dynamics and Control
At the Center for Computational Quantum Physics, we are developing the conceptual basis, theoretical formalism and computational tools needed to use the quantum nature of light to understand and control quantum phenomena in complex systems.
CCQ
Exoplanets
Exoplanet researchers at the Center for Computational Astrophysics (CCA) study the origins and evolution of planetary systems around other stars, from simulations of their initial formation to observations of their present-day conditions.
CCA
Galactic Chemo-Dynamics
Galaxy Formation
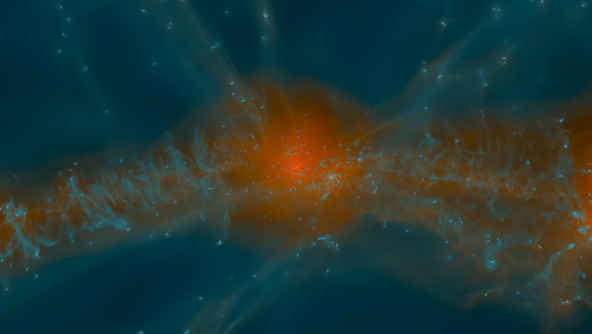
The Galaxy Formation group is developing the numerical tools and physical insights necessary to understand how galaxies form and evolve within a cosmological context.
CCA
Genomics
An immensely complex molecular network of interactions forms the foundation of human biology and disease. Genomic approaches provide a particularly illuminating window to biological systems, and when combined with advanced analysis allow us to learn and model this complexity.
CCB
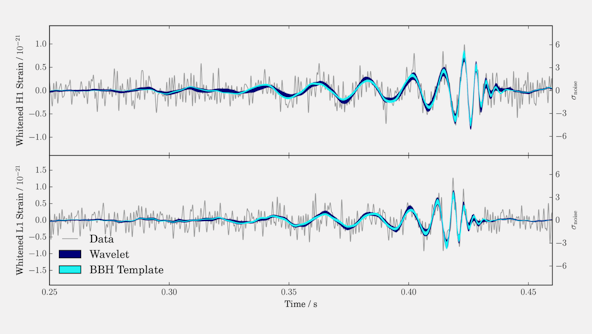
Gravitational Wave Astronomy
The detection of gravitational waves from the merging binary black hole GW150914, followed by the (likely) binary black hole mergers LVT151012, GW151226, and GW170104, means the era of gravitational wave astronomy is upon us.
CCA
Image and Signal Processing
At CCM, we develop new computational and statistical methods, and accompanying software, for the analysis of large, complex data sets, especially those that arise from high-throughput scientific experiments.
CCM
Initiative for Computational Catalysis
The Simons Foundation Initiative for Computational Catalysis is a new effort within the Flatiron Institute dedicated to the development and application of computational techniques enabling the rational design of more efficient and more affordable catalysts.
Interstellar Medium
Machine Learning X Astrophysics
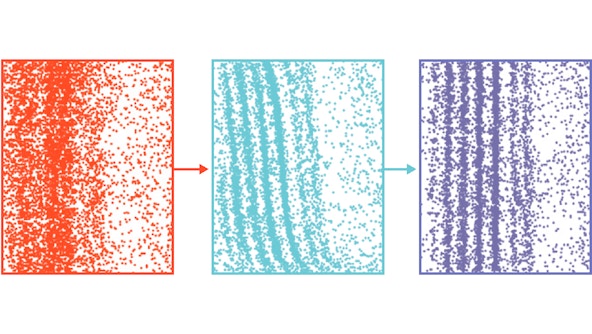
Machine learning is being increasingly integrated into scientific discovery to augment and accelerate research, helping scientists to generate hypotheses, design experiments, collect and interpret large datasets, and gain insights that might not have been possible using traditional scientific methods alone. Researchers at CCA is leading the wave of rapid development and adoption of machine learning techniques to enable and accelerate scientific discovery.
CCA
Nearby Universe
The Nearby Universe group studies the structure, growth and dynamical evolution of our galaxy and its neighbors. We use the nearby universe as a laboratory to constrain dynamical processes and the properties of dark matter. The group works at the interface between large observed and simulated datasets.
CCA
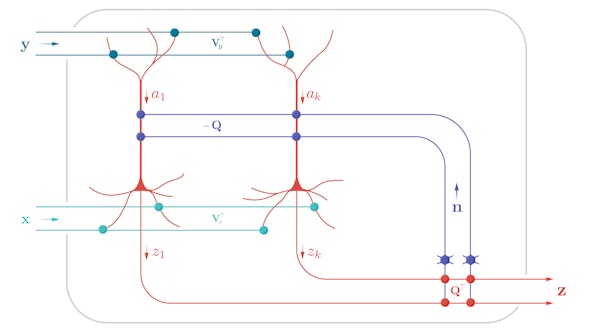
Neural Circuits and Algorithms
Our goal is to understand how the brain analyzes large and complex datasets streamed by sensory organs in order to aid efforts at building artificial neural systems and treating mental illness.
CCN
NeuroAI and Geometric Data Analysis
Our group develops mathematical theories for understanding how neurons collectively give rise to behavior in biological and artificial neural networks. Our current focus is on addressing this question through two broad approaches at the intersection of computational neuroscience and deep learning:
CCN
Numerical Analysis
The direct numerical simulation of many scientific processes remains impractical, even with modern supercomputers.
CCM
Particle Astrophysics
CDS
Planet Formation
Plasma Astrophysics
Polymathic AI
Quantum Materials
What arrangement produces a material with an optimal thermoelectric figure of merit, or a combination of magnetic and ferroelectric properties, or a high superconducting critical temperature?
CCQ
Resolved Stellar Populations
Software Libraries
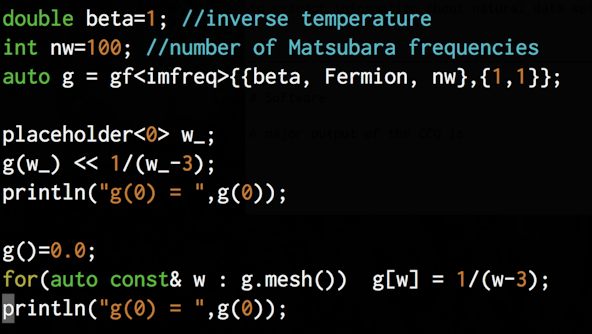
A major effort of the CCQ is the development and support of high quality open-source software for quantum many-body physics research. Making software which is reliable, efficient, and productive accelerates discovery and fosters scientific consensus and reproducibility.
CCQ
Stars
CO
Stars and Compact Objects
The Stars and Compact Objects group develops the theoretical framework and computational methods needed to study the life and death of stars, as well as the astrophysics of neutron stars and black holes.
CCA
Statistical Analysis of Neural Data
We develop statistical models and open-source computational tools to extract insights from neural data. We are particularly interested in characterizing flexibility and variability in neural circuits—e.g., how do the dynamics of large neural ensembles change over the course of learning a new skill, during periods of high attention or task engagement, or during development and aging.
CCN
Structural and Molecular Biophysics
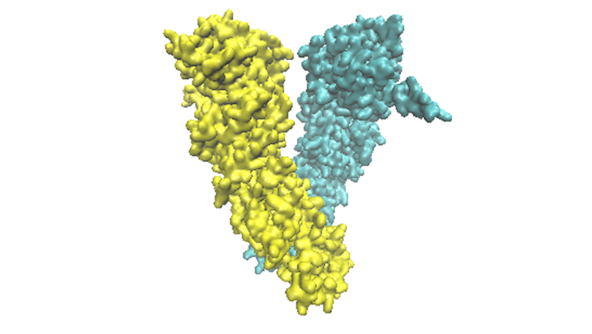
Underlying all biological processes are molecules and their interactions with each other. However, our ability to understand how these molecules function over biologically relevant scales remains very limited.
CCB
Theory and Methods
The quantum many-body problem is one of the hard problems of complexity theory: A direct solution requires computational resources that grow exponentially with the size of the problem to be solved.
CCQ
Advancing Research in Basic Science and MathematicsSubscribe to Flatiron Institute announcements and other foundation updates