[Updated April 12, 2017, with availability information, below.]
Complete genome sequences from more than one hundred diverse human populations
The largest dataset of diverse, high quality human genome sequences ever reported is presented below.
The sampling strategy differs from studies of human genome diversity that have aimed to maximize medical relevance by studying populations with large numbers of present-day people. This new study takes a different approach by sampling populations in a way that represents as much anthropological, linguistic and cultural diversity as possible, and thus includes many deeply divergent human populations that are not well represented in other datasets.
All genomes in the dataset were sequenced to at least 30x coverage using Illumina technology. The sequencing reads were mapped and genotyped using a customized procedure that was optimized for population genetic analysis. The researchers eliminated bias of alleles toward matching the human genome reference sequence, and determined genotypes on a single-sample basis to avoid preferential calling of genotypes from populations that had more individuals represented.
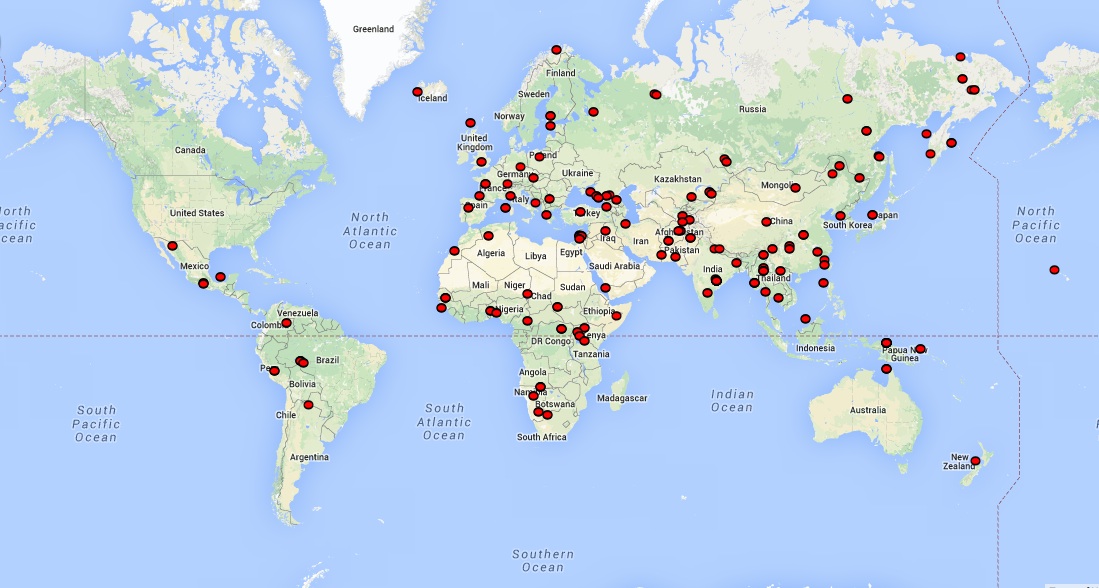
Metadata on the samples can be found here.
The primary dataset (Panel C in the first column of the metadata file) consists of data from 260 genomes from 127 populations: 39 Africans, 23 Native Americans, 27 Central Asians or Siberians, 49 East Asians, 27 Oceanians, 38 South Asians and 71 West Eurasians. For convenience, genotyping results for an additional 18 genome sequences published previously are also included.
The data include Variant Call Formats files (VCFs) with genotype calls at every position in the genome. The consortium also plans to release BAM files containing the raw sequencing reads.
A README containing directions for downloading can be found here.
Please note that there are approximately 10 terabytes of data and because of the large dataset size, the data need to be downloaded using the gridFTP software after applying for and obtaining a certificate from the hosting site.
Questions
If you have questions about the dataset, please contact Shop Mallick ([email protected]), Nick Patterson ([email protected]) or David Reich ([email protected]). If you have problems with the dataset once you download it, we would be grateful if you could let us know so that we can fix any issues in future releases.
Use of the genome sequence data (Please respect Fort Lauderdale principles)
All data are made freely available. However, please observe the Fort Lauderdale principles, which entitle the data producers to make the first presentation and publish the first genome-wide analysis of the data. The data can be used freely for studies of individual genes or other individual features of the genome.
Information on an earlier set of genomes released from the same set of researchers, ‘A publicly downloadable dataset of 25 deep genome sequences of which 13 are experimentally phased,’ can be found here.
Update: April 12, 2017
Simons Genome Diversity Project dataset is now available on the Seven Bridges Cancer Genomics Cloud
Data Portal
Access the project on the Seven Bridges Cancer Genomics Cloud (CGC) through the data portal. After you log in to the CGC with a free CGC account or with existing credentials, you will be taken to the Simons Genome Diversity Project (SGDP) dataset and public project.
About the Portal
The Cancer Genomics Cloud (CGC), powered by Seven Bridges, is one of the three pilot systems funded by the National Cancer Institute to explore the paradigm of colocalizing massive genomics datasets alongside secure and scalable computational resources for analyzing them.
Seven Bridges recently ported the complete, open-access SGDP dataset, version-matched with the latest SGDP publication in Nature, to the CGC. The dataset contains 279 complete genome sequences from 130 diverse human populations. It is organized as a public project, where you can filter and query files of interest and use Common Workflow Language analysis tools to interrogate the genomics landscape of different populations. You will not be charged for the storage of any SGDP files.
Learn more about using the SGDP public project.