Spike Sorting Becomes Fully Automated

New technologies allow neuroscientists to record the activity of hundreds to thousands of cells. But sorting through all those data — a mess of electrical spikes from many different cells — is a challenge. To make sense of it, researchers need to figure out which spike came from which cell, often a prohibitively laborious process.
More and more neuroscientists are reaching the point where they can no longer do this kind of analysis by hand. “Automating spike sorting is something we absolutely have to have for the next generation of neuroscience studies,” says Jonathan Pillow, a neuroscientist at Princeton University and an investigator with the Simons Collaboration on the Global Brain (SCGB). “With these large-scale datasets, it will not be feasible for humans to do spike sorting anymore.”
A new, fully automated method to sort spikes, published in Neuron in September, could help solve this problem. The software, called MountainSort, was developed at the Simon Foundation’s Flatiron Institute, in close collaboration with Loren Frank, a neuroscientist at the University of California, San Francisco and an SCGB investigator. “The objective is to get humans out of the equation as much as possible,” Frank says. Researchers have made MountainSort freely available on GitHub.
“For this application, we think it’s better than any of the other algorithms,” says Jeremy Magland, a data scientist at the Flatiron’s Center for Computational Biology who developed MountainSort.
Frank says the technology has been transformative in his own lab, which is working on high-density recordings with polymer electrodes. “We have been able to sort data that would have taken years in days to weeks,” he says. “You can use it with little or no human intervention, which is the main problem with other spike-sorting algorithms.”
Elizabeth Buffalo, a neuroscientist at the University of Washington in Seattle, and an early adopter of MountainSort, concurs. Her lab recently started using arrays with 124 electrodes, making manual sorting untenable. “We were really facing a bottleneck of not being able to analyze our data,” says Buffalo, an SCGB investigator. “We just started using MountainSort, but it feels like a game changer for our lab.”
Multi-electrode arrays are made up of tens — sometimes hundreds — of electrodes, all recording neural activity from overlapping subsets of cells. Multiple electrodes detect each spike, creating a complex jumble of electrical signals. Scientists typically attribute specific spikes to individual cells based on the shape of the voltage trace, or waveform. However, neurons with similar locations and geometries can produce nearly identical electrical responses, making it difficult to distinguish between them.
Humans sort spikes by clustering like spikes together. Researchers plot each spike in two-dimensional space according to different variables, such as the amplitude of the waveform. Waveforms from the same cell will cluster together, providing a clear way to group activity from individual cells. But the process is laborious.
Spike-sorting algorithms take the same approach on a much larger scale. However, most require human intervention to make sure the clustering is accurate. Clusters with certain patterns, such as a non-Gaussian distribution, can prove troublesome. “One of the challenges with spike sorting is that signals are complicated and not easy to describe mathematically,” Frank says.
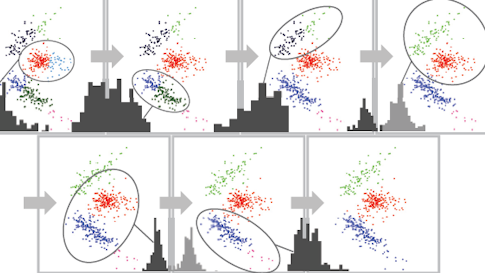
MountainSort removes the human-review step by employing an algorithm that makes few assumptions about the data it processes. The program starts by over-sorting spike data into many small clusters. Then, it iteratively evaluates whether neighboring clusters should be joined, by looking at the density of data points in those pairs of clusters.
Since the paper was published in early September, Magland has received daily inquiries about the software. He estimates that roughly a dozen labs are starting to use the software, with 10 to 20 more considering it.
In addition to speeding up spike sorting, researchers say that MountainSort will generate more reproducible data, because it removes the subjectivity that comes with manual sorting. “You can share data because it’s not subject to a user’s arbitrary decisions,” Frank says. Users can annotate the software’s clusters, but the original clustering is preserved, allowing outside researchers to evaluate the quality of the data.
“We really like the motivation of having something completely automated; I think that’s going to enhance rigor and reproducibility across labs,” Buffalo says. “It’s harder to take the dataset at face value if you don’t know the quality of sorting that went into it.”
One challenge the researchers are still working on is how to deal with electrodes that drift in the brain over time. “In long recordings, the shapes of the waveforms change due to slight movements of the implanted electrode or other factors,” Magland says. In these cases, the algorithm may mistakenly categorize spikes over time as coming from two different cells. Magland and his colleagues are working to develop techniques to overcome this and other challenges.


