Computation Opens New Doors for Science

Many areas of scientific research today would not exist at all without computer science. “Fifty years ago, you did a scientific experiment, and you extracted a number or a measurement,” says Leslie Greengard, a physician, computer scientist and acting director of the Flatiron Institute, the research division of the nonprofit Simons Foundation in New York City. “Now, because of certain kinds of automation, it’s possible to spew out just reams of data. And it’s useless as is. The mapping from the data to something that’s scientifically interpretable involves a huge number of steps that people don’t really appreciate. And all those steps are computational.”
In other words, computer programs can process that abundance of data to tell us the structure of a protein or a galaxy or a brain. Yet when it comes to the relationship between scientists and programming, well, it’s complicated.
The academic community has never been great at building and maintaining software tools. Most software projects have user bases that are too small to garner commercial interest, but the projects are too large for a single lab to handle. “I basically took this job because I’ve always found that gap very unfortunate,” says Greengard, who joined the foundation in 2013 and helped create the Flatiron.
“The culture of software development in the academic world is a challenging one,” says Nick Carriero, a computer scientist at the Flatiron. “People who are coming up as computational biologists or computational astrophysicists or computational neuroscientists are spending a lot of time learning their science. And it’s often kind of expected that they pick up the computing as they go along. But there isn’t a significant commitment to formal training on the computing side.”
The result? The software critical to advancing the science can sometimes suffer. Moreover, the graduate students who develop the code graduate, moving on to other labs, making it difficult for new students to easily adopt the work and keep it updated. “What do you do if you build increasingly complicated software packages that need some care and feeding, but it’s not naturally anyone’s responsibility?” Carriero says. “Recognizing that dilemma was part of the origin story for us, and for the Flatiron as a whole.”
By “us,” Carriero means the Scientific Computing Core (SCC), which runs the Flatiron’s computational infrastructure. Carriero and Ian Fisk, a trained physicist, direct the SCC, a resource unlike any other. The SCC employs a team of 14 experts in various aspects of computing in an institute of about 200 and plays several roles. The team builds and maintains computing and network hardware, writes software code, and collaborates with scientists to optimize computational tools. Universities have IT departments, Greengard says, but such departments are not involved in the research. National labs have people who play roles similar to that of the SCC team, “but even there,” he says, “it’s rare to find things at this scale.”

Open-door policy
The Simons Foundation was co-founded in 1994 by Jim and Marilyn Simons, and for 20 years it was dedicated to supporting research in mathematics and the basic sciences by making grants to investigators and projects at academic institutions. In 2013, the Simonses began to make room for research in-house, focusing on data-intensive biology. They created the Simons Center for Data Analysis (SCDA) and tapped Greengard to helm it. Greengard then hired Fisk and Carriero, who started on the same day, and the group grew to 50 people. Over the next few years, the SCDA evolved into the Center for Computational Biology (CCB) and was joined by the Center for Computational Astrophysics (CCA) and the SCC. The Flatiron Institute was created in 2016 to house the centers, which were later joined by the Center for Computational Quantum Physics (CCQ), the Center for Computational Mathematics (CCM) and, most recently, the Center for Computational Neuroscience (CCN).
Meanwhile, computing needs expanded. “I used to joke that some strategy meetings were held in my shower because there was only just me,” Fisk says. The SCC’s original computing cluster consisted of Macs stacked on employees’ desks. Today, 100,000 dedicated cores are spread across four locations: the Flatiron Institute; Brookhaven National Laboratory on Long Island; Secaucus, New Jersey; and the San Diego Supercomputer Center. The network connecting the computers started with the bandwidth of a home user but has grown by a factor of 10,000.
But the hardware is only half the story. The SCC is a hands-on outfit. In some cases, they dedicate someone to help with a scientific project. Other times they offer advice to scientists who get stuck with their code. Or, after observing the behavior of the clusters, they might spot software anomalies and reach out. “We might proactively go to the researcher,” Carriero explains, “and say, ‘You know, it looks like your code is doing something a little strange. Why don’t we sit down and talk about what you’re doing? And maybe we can make some modifications, which will both make the machines run more efficiently, and get you your answer a little bit more quickly.’” Carriero emphasizes that the SCC is sensitive to scientific context. “So even though my background is in computer science, and we tend to chase after the latest developments in programming languages, say, we will not tell someone who walks into our office who is needing help, ‘The first thing you have to do is rewrite your entire code into the latest language.’” Instead, they’ll suggest more modest modifications.
The SCC is the only Flatiron division with staff on every floor of the institute. “It’s meant to break down a wall that you often see in the academic research computing world,” Carriero says.
Most of the software they develop is open-source and available for anyone in the scientific community or industry. The SCC also conducts tutorials and creates publicly available videos that introduce basic software development principles, such as testing, loading modules, Linux and version control.
Mapping genes and brains
The SCC helps with a variety of projects across the five Flatiron centers. One of these tasks at the Center for Computational Biology and the Center for Computational Mathematics is to determine protein structure. In the last decade, biologists have adopted a new method called cryogenic electron microscopy, or cryo-EM. They spray a layer of proteins across a microscopy slide, flash-freeze it, and take images. Then they use computers to reconstruct the protein from the points of view of many protein copies lying at different angles. Fisk compares it to constructing a 3D model of a face from multiple photos taken at different angles. Cryo-EM, whose developers won a Nobel Prize in chemistry in 2017, is revolutionary in its importance for drug development, especially for those treating complex diseases of the brain. And this groundbreaking work would not be possible without computational power provided by research computing centers like the SCC.
Another biological endeavor, outside the Flatiron but within the foundation, is SFARI, the Simons Foundation Autism Research Initiative. In addition to granting tens of millions of dollars in research funding annually, SFARI supports hundreds of labs by developing and maintaining genetic databases, animal models, brain tissue samples, cell lines and data analysis tools. A central focus is mapping the genomes of families touched by autism. But each person’s genome consists of 3 billion base pairs; in computing terms, this translates to a 200-gigabyte file per individual. To reach statistical conclusions about genes relevant to autism, researchers must compare genomic data from many individuals. Just obtaining and sharing the data — totaling 2 petabytes for 10,000 people — is a challenge that requires the SCC’s help. In its early days, Fisk would retrieve the files from the sequencing centers on hard drives via taxi. (“On the way back, it was probably the fastest network in all of New York,” he says.) Soon, the SCC installed higher-capacity internal networks and external network access. Still, getting the files to the 200 labs worldwide that use them was another matter. The SCC developed an access portal for SFARI and modified the networking software so labs could download large files more efficiently. “Basically, what it tries to do is keep relatively large network pipes busy,” Fisk says. Natalia Volfovsky, director of data and analytics at the foundation, says numerous papers are published every year using the data.
The Center for Computational Neuroscience recently emerged as its own center led by Eero Simoncelli, having begun under the auspices of the CCB. One of its first major projects will be to build a connectome, a wiring diagram of an entire brain that accounts for all its neurons and the connections between them. Creating connectomes is a long-standing scientific challenge made difficult by the unparalleled complexity and messiness of the densely packed brain. The first connectome, of a roundworm with just 302 neurons, took scientists many years to complete.
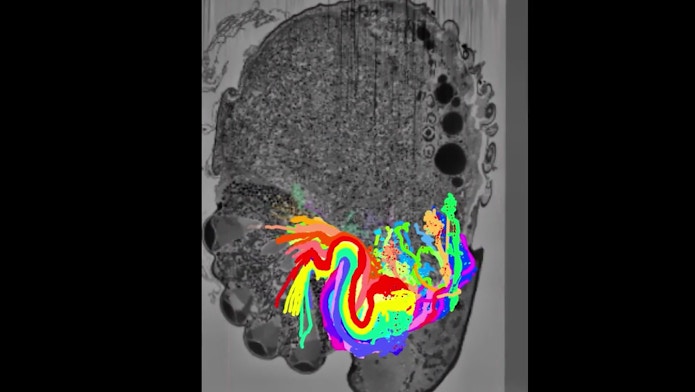
Dmitri “Mitya” Chklovskii, a neuroscientist and group leader at the CCN, says he and his team are mapping a microscopic wasp brain, which could reveal critical clues to how larger brains function. Indeed, the wasp performs many of the same functions as larger, more complex animals — it flies, sees and smells — but its brain has only 7,400 neurons, far fewer than the human brain, with its about 86 billion. To accomplish this, Chklovskii’s collaborators sequentially ablate the brain in eight-nanometer steps while photographing each section using electron microscopy. From millions of such photographs, scientists reconstruct the 3D shapes of individual neurons. “This is like, you had a bowl of spaghetti, you froze it, and you sliced it up, and then you want to trace the outline of each noodle,” Chklovskii says. Working manually, scientists first outline the neurons and their tendrils in many of the images. Then, Chklovskii’s team trains computers on the hand-labeled images to automate the process and merge the data into 3D models. The microscopic wasp dataset is not huge, about 1 terabyte, but the computer labeling requires extreme precision. “That goal is very ambitious.” Creating connectomes like this is expected to help understand behavior. “If you could decode some ancient language in which nature wrote the brain,” Chklovskii says, you could read memories, treat disease and maybe someday build better artificial intelligence.
By clicking to watch this video, you agree to our privacy policy.
Pat Gunn, an SCC software engineer, is embedded in Chklovskii’s group about half the time. One problem he helped solve was combining software that was not designed to work together — “It’s like a zoo of separate pipelines,” Gunn says — allowing researchers to, for instance, visualize the computer’s output while editing its mistakes. Training the machine learning software is also computationally intensive, and Gunn has helped get it running on the San Diego cluster. “If Mitya is a CEO of a research project,” Gunn says, “then I get to kind of be the CTO. And so that’s fun.” Gunn studied computer science but has taken courses in neuroscience, which come in handy. “I can understand the challenges both on a scientific level and on a technology level.”
Gunn has also helped Chklovskii’s group with another project involving software called CaImAn. Neuroscientists can introduce molecules into animal brains that glow in the presence of calcium, serving as a marker for neural activity. “This gives you an ability to watch the work of the brain in real time,” Chklovskii says. CaImAn translates videos showing that activity into data on the activity of individual neurons. “The SCC supported this project to the point where our package is now one of the top packages for calcium imaging in the community,” Chklovskii says. “There are hundreds of labs that use this.”

Miniature universes
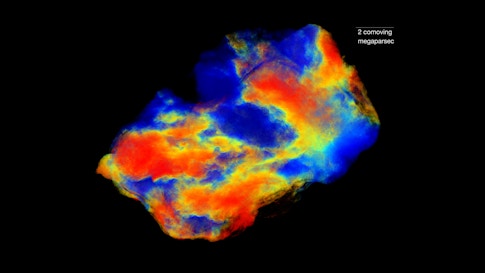
Far afield from biology, the Center for Computational Astrophysics relies on the clusters to run large simulations. Shy Genel, a member of the CCA, studies galaxy formation, hoping to understand what determines galactic rotation rates, why galaxies tend to be of two main types, and what they can tell us about the universe’s contents. “This is a field that’s really, really demanding in terms of the computation power,” he says. Instead of using telescopes, he builds mini-universes containing hundreds or millions of galaxies, millions of light-years across, and runs them from almost the beginning of time, 14 billion years ago. “In terms of the support that I’m getting from the SCC,” he says, “on the most basic level, we have a really big cluster here, and it runs just really amazingly smoothly. Hiccups are extremely rare.” He’s worked at Harvard, Columbia, the Max Planck Institute and elsewhere, he says, but nothing compares.
“And then at a deeper level, this team is really working with you, for you,” Genel says, in ways uncommon elsewhere. The SCC team looks into his code to see how it could run faster, diagnoses deep problems, and even supports his students who have less computing experience. One key to accelerating the code was to improve its ability to break up a task into pieces that could run on thousands of CPUs in parallel. This reduces the time it takes a simulation to run from years to months. Genel is now exploring how machine learning can speed up the process even further. He and SCC have run 2,000 simulations with slightly different laws of physics, and hope to train AI to approximate the same results in less time, so that future simulations can use AI approximations. The dataset of simulations is public so others can do the same.
Genel hopes to challenge the field of computational astrophysics to find ways to combine two types of expertise. “It’s hard to imagine an astrophysicist just describing what they need and handing over some definition document to a software engineer, and at the same time you’re not going to have enough software engineering training for astrophysicists,” he says. “So I think as code gets more complicated and computer clusters gets more complicated, it’s probably getting more and more urgent to find ways to really work together and develop efficient, good code for simulating physical systems.”
Strange states of matter
The Center for Computational Quantum Physics also simulates miniature universes, but at a much smaller scale. Nils Wentzell, a CCQ data scientist, joined the Flatiron to continue developing the software package TRIQS, or Toolbox for Research on Interacting Quantum Systems. If you want to simulate the behavior of electrons in a periodic lattice of atomic nuclei, you need to approximate their interactions because of the sheer number of particles. The TRIQS software provides all the necessary tools to implement such approximae schemes on the computer. Overall, Wentzell says, CCQ studies interesting phases of matter linked to the interactions of electrons — not typical solids, liquids and gases, but things like superconductors, in which electrical current can pass without resistance. Typically, superconductivity requires very low temperatures, but materials that superconduct at room temperature could revolutionize power grids and technologies that rely on strong magnets, such as medical imaging, levitating trains and fusion reactors.
TRIQS can run on a personal computer, but large calculations require high-performance computing clusters, which is where the SCC comes in. Wentzell says the SCC has made the software not only scalable, but also easier to install, run and test, by bundling it with other software libraries it depends on. The SCC team has also helped in training new users.
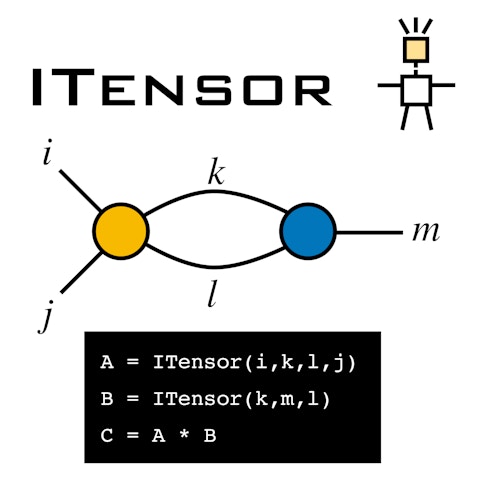
Miles Stoudenmire, another CCQ scientist, is developing a software package called ITensor, which simplifies the manipulation of large, multidimensional arrays of numbers. Dylan Simon, a member of the SCC, has helped the code run more smoothly and generate performance reports. “He’s like a wizard at that stuff,” Stoudenmire says. “That may sound a little bit trivial, but it’s one of those day-to-day things that we’re using constantly; it raises our code quality and our ability to keep track of what we’re doing.”
Tinkering with black boxes
While some groups use neural networks, a form of artificial intelligence, as tools to understand brains or galaxies, some members of the Center for Computational Mathematics try to understand the networks themselves, to open the “black boxes” that magically translate questions into answers. Greengard, who started at the CCB, now heads the CCM. He and his colleagues spend their days inventing state-of-the-art algorithms (not necessarily neural networks). “It’s sort of like doing plumbing,” he says. “I generally don’t work on the applications themselves. My driving interest generally isn’t a deeper understanding of electromagnetics or acoustics or fluid dynamics. I just like creating solvers that let other people do such simulations effectively.” The work sometimes ends up in academic publications and sometimes in tools for the other centers and for scientists outside the Flatiron. Greengard sees his work and the SCC’s as complementary. “What the SCC has done extremely effectively is to look at the computation by scientists in the other centers and make recommendations that improve the algorithmic speed,” he says. “But the SCC folks, by and large, don’t say, ‘Let me step away from this problem for two years and see if I can come up with a better approach for solving that class of problems.’” That’s Greengard’s department.
While Greengard’s projects sometimes start out very theoretical, he eventually wants the products he develops to work in high-performance computing environments. His current projects are still immature, but in the coming years, “I’m actually very interested in making that transition” with the help of the SCC, he says. “I’m not sure they know about that, exactly. But they’ll hear soon.”


