aLENS
View Project
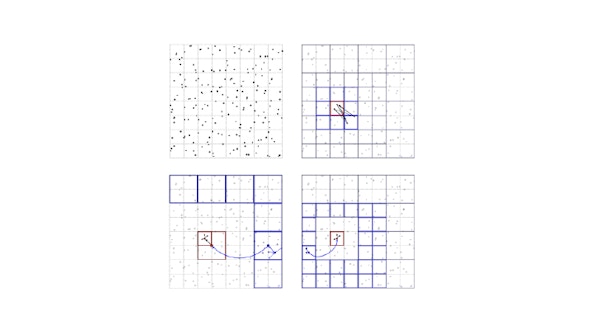
This is the simulation tool for tracking assemblies of microtubules driven by motor proteins. Microtubules are modeled as straight rigid fibers and motor proteins are modeled as Hookean-springs with active binding-unbinding dynamics governed by a two-stage kinetic Monte-Carlo model to impose detailed balance. A state-of-the-art collision resolution algorithm based on geometrically constrained optimization is incorporated to reach long physical dynamics timescales. This package is massively parallel.