CaImAn Python
View Project
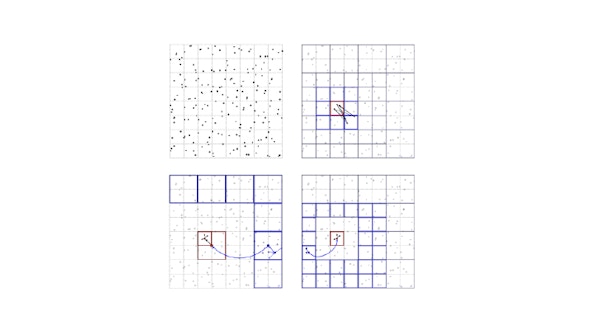
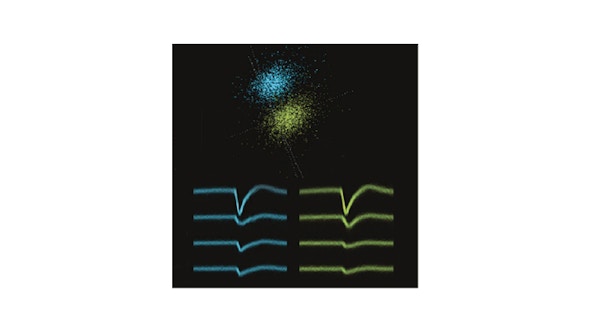
Recent advances in calcium imaging acquisition techniques are creating datasets of the order of Terabytes/week. Memory and computationally efficient algorithms are required to analyze in reasonable amount of time terabytes of data. This project implements a set of essential methods required in the calcium imaging movies analysis pipeline. Fast and scalable algorithms are implemented for motion correction, movie manipulation, and source and spike extraction. CaImAn also contains some routines for the analyisis of behavior from video cameras. In summary, CaImAn provides a general purpose tool to handle large movies, with special emphasis on tools for two-photon and one-photon calcium imaging and behavioral datasets.