Introducing ROCKET: An Upgrade to AlphaFold That Learns From Raw Experimental Data

A new software tool called ROCKET is giving the Nobel Prize-winning AI-powered AlphaFold a serious upgrade in its ability to predict how proteins fold.
Proteins are the workhorses of our cells, and how they fold up like complex origami determines their behavior. When this folding goes awry, it can lead to conditions such as neurodegeneration and cancer. Predicting how proteins fold was long considered one of the greatest challenges in biology.
AlphaFold revolutionized the field by using machine learning to predict protein folding based on training data. But despite the huge amount of new raw data that has become available since AlphaFold’s launch, the software isn’t equipped to directly interact with it and learn from it. AlphaFold can only accommodate atomic models of the data — representations that make the data more interpretable, but that can take weeks of human effort to build.
ROCKET extends AlphaFold’s capabilities to accommodate raw experimental data. Its creators present the tool in a paper published April 1 in Nature Methods, demonstrating its ability to handle model building from data sources such as X-ray crystallography and cryo-electron microscopy experiments in a fully automated and reliable manner.
The upgrade opens the door for a better understanding of how proteins change shape as they interact with other proteins, hormones, metabolites and their physical environment — information critical to designing new drugs to fight diseases.
“This is exciting work that allows powerful protein structure prediction models like AlphaFold to interface directly with raw experimental measurements,” says Minhuan Li, a Flatiron Research Fellow at the Flatiron Institute’s Center for Computational Mathematics and a former graduate student in Doeke Hekstra’s lab at Harvard University. “Automating this step makes it much easier to process large, high-throughput datasets.”
Li serves as co-lead author of the new paper with Alisia Fadini of the University of Cambridge. Li and Fadini worked on the project with collaborators from Columbia University, Harvard University, Umeå University, Meijo University, Trinity College Dublin, St. Jude Children’s Research Hospital and the Karolinska Institute.
Inside ROCKET
“We got very excited about the possibility of using all the knowledge baked into models like AlphaFold to help us interpret our experimental data,” says Hekstra, one of the study’s senior authors.
As a first step, Li built SFCalculator, a tool that links atomic models to their underlying data.
“SFCalculator provides a fast, differentiable way to simulate experimental observations from atomic models,” says Li. “It creates a seamless bridge between raw data and modern AI, which allows powerful, pretrained deep learning models to be directly guided by experimental results.”
Li and Hekstra then began speaking with Fadini, a postdoc working with Randy Read and Airlie McCoy at the University of Cambridge, and Mohammed AlQuraishi at Columbia University. They realized they were on the same path. AlQuraishi’s group had created ROCKET’s other building block, OpenFold, an open-source version of AlphaFold. Li and Fadini integrated SFCalculator with OpenFold and voila: ROCKET was born.
“Once Alisia and Minhuan put these pieces together, their progress was breathtaking,” says AlQuraishi.
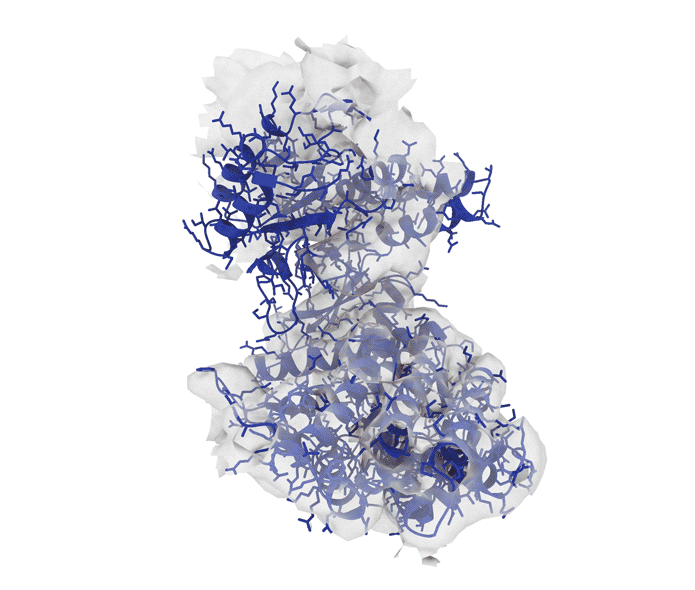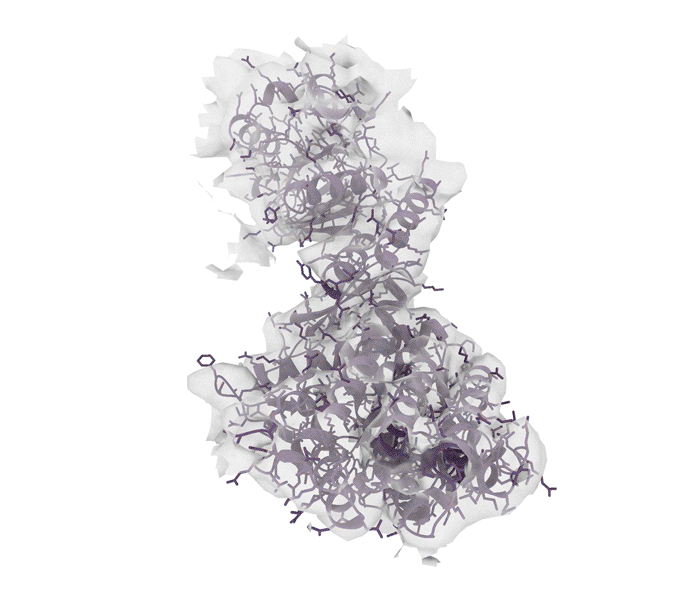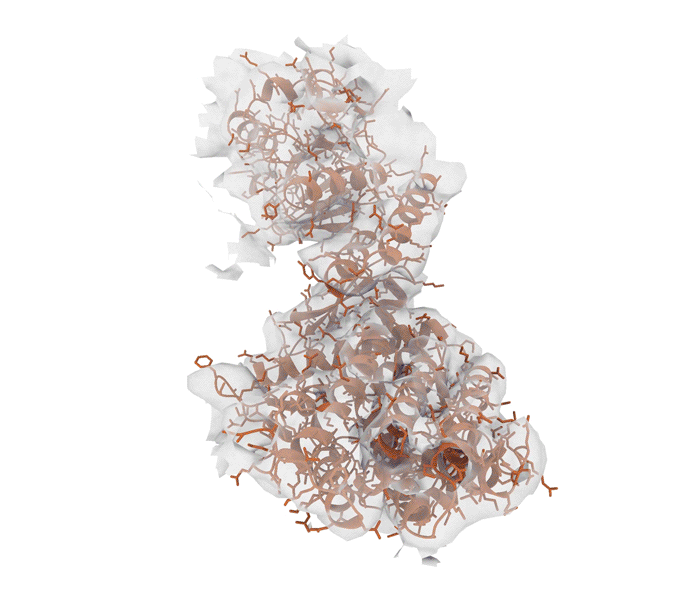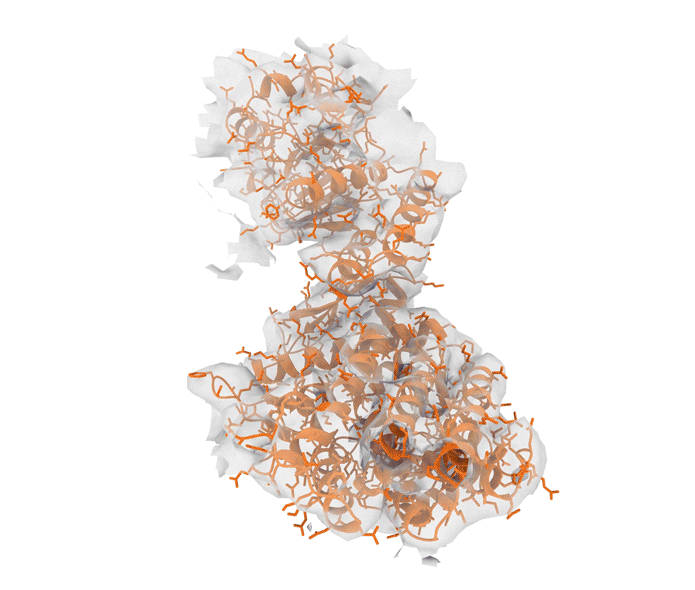
Excitingly, ROCKET can handle structural data from advanced imaging tools such as X-ray crystallography (which determines the atomic and molecular structure of crystals) as well as cryo-electron microscopy (cryo-EM) and electron tomography (ET), which produce detailed 3D models of proteins, viruses and cells.
“Although we started with X-ray crystallography, it has been stunning to see just how well the approach extended to cryo-electron microscopy and tomography,” says Hekstra. “We see that the prior information encoded by AlphaFold is especially helpful in interpreting low-resolution cryo-EM and ET data. That’s where ROCKET really shines.”
Scientists already knew that examining how protein sequences have changed over evolution can provide clues about the different shapes a protein might adopt. But it wasn’t clear how to get AlphaFold to leverage those clues to accurately predict multiple possible shapes. ROCKET, however, is good at this.
“We found that ROCKET can jump from one plausible conformation to another,” says Hekstra. “It is quite striking to watch ROCKET as it searches: The jumps it makes are unlike anything traditional methods would do.”
The team recently hosted a webinar on ROCKET to demonstrate its use, and will soon make the tool openly available to the research community. The tool will be integrated into PHENIX (the widely used structure-determination software platform), Harvard’s SBGrid (a platform that delivers ready-to-use structural biology tools to academic and industry researchers around the world) and RS-Station (an open-source software development platform and community that originated in the Hekstra lab).
“Handling the massive scale of today’s high-throughput experimental data requires robust, open-source infrastructure,” says Li. “It takes a tremendous amount of effort to translate a theoretical research concept into a practical tool — one that researchers can seamlessly drop into their existing pipelines — but it is a critical step. Our ultimate goal is to ensure these tools are genuinely accessible and useful to the community.”