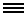A computer simulation of a network of microtubules that has contracted due to the pulling action of dynein motor proteins traveling along them. The collapsing network generates material stresses thought to be important to the shape of many mitotic spindles. This simulation is by Flatiron research fellow Wen Yan and staff scientist Sebastian Fürthauer of the CCB’s biophysical modeling group.
The DNA inside a cell’s chromosomes contains a blueprint for assembling and regulating all the cell’s proteins, but that blueprint is not simply lying open, waiting to be read. Instead, tiny ‘machines’ within the cell are constantly thumbing through the pages, moving the fibers of DNA to bring certain regions together into useful combinations. And when a cell divides, the chromosomes get moved about even more: A spindle structure pushes and pulls them into formation and escorts them into the newly born cells.
“All of this movement factors into how cells work and how they divide,” says Michael Shelley, leader of the biophysical modeling group at the Center for Computational Biology in the Flatiron Institute, a
division of the Simons Foundation. “It’s not just a matter of molecules and chemical reactions — there’s physics in there, and fluid dynamics too.”
Shelley and his collaborators are developing mathematical models to describe how spindles form and chromosomes move. Their work is part of an emerging field called ‘active matter’ that studies, as Shelley puts it, “how biological systems put themselves together.” Unifying soft condensed matter physics with the life sciences, active-matter problems “go right to the heart of fundamental aspects of biology,” says Shelley, who is also an applied mathematician at New York University.
The centrosomal arrays of nuclear complexes are involved in the complex’s positioning prior to cell division, but exactly how this happens is unclear. This simulation investigates one model of the process, wherein microtubules in the array grow and push against the cell wall before depolymerizing, and in this way slowly center and rotate the complex into its proper place and orientation. The shapes of the dynamical microtubules and the flows of cytoplasm within the cell are signatures of particular positioning models. These shapes can be compared with the results of an experiment to help identify how forces are transduced within the cell to accomplish this crucial function. Work by Ehssan Nazockdast and Michael Shelley of CCB
Active matter focuses on systems — spindles in a cell, swarms of bacteria, schools of fish or flocks of birds — made up of many individual components that collectively generate motion or mechanical stresses. Finding the equations governing these collective behaviors could bring researchers closer to a quantitative theory of some of life’s central phenomena.
Doing so will require a deep understanding of the role fluids play in such systems, Shelley says. “The fluid is what lets individuals in a system ‘talk’ to each other remotely,” he says. “Individuals may run into or push each other, but the fluid they move can organize them collectively.”
Fluid flow is a driving force in both microscopic systems, such as cellular spindles, and macroscopic systems, such as schools of fish. Yet fluid dynamics also poses a special challenge when it comes to establishing a unified theory of such systems. In microscopic systems, the fluid communicates the motion of one component across the system almost instantly. If the individual component stops moving, the fluid also stops moving and communication ceases. But in the water and air through which schools of fish and flocks of birds move, information can live on, like the vortex trails left by an airplane, even if the components stop moving. Reconciling the immediate loss of information in some systems with its lingering effects in others will be a challenge, Shelley says.
How biopolymers and motor proteins interact is important to the creation of new active materials and to achieving a basic understanding of how cellular organelles, such as spindles and cortices, assemble and regulate themselves. This simulation shows how the bundling by dynein motor proteins of microtubule minus-ends can lead to the formation of a contracting material network. The measured degree and speed of contraction is consistent with experiments using cellular extracts. In addition, measured stresses on material support an ‘active-materials’ continuum model of mitotic spindle assembly. Work by Wen Yan, Sebastian Fürthauer and Michael Shelley of CCB
For now, Shelley’s group is focusing on developing computer models and simulations of the interactions between a cell’s fluid cytoplasm, its spindles and other cellular structures. “In this cellular-mechanics world, we’re doing a level of computational fluid dynamics that no one else does,” Shelley says. He hopes his team’s numerical and computational tools will allow researchers to work on other problems in active matter.
“It’s going to be exciting figuring out how we can interface with the other groups in the center,” Shelley says. “It’s going to push all of us into thinking about problems we might not ever consider otherwise.”