Neuropixels Offer New Ways To Map Cells
On a chilly evening in November 2017, neuroscientists packed a conference hall for a last-minute session at the annual Society for Neuroscience meeting in Washington, D.C. The crowd was eager to hear about Neuropixels, a novel technology described in Nature that month that enables scientists to simultaneously record from hundreds to thousands of cells across different regions of the brain. (For an introduction to Neuropixels technology, see “‘Neuropixels’ Expand Access to the Brain” and “Coming Soon: Routine Recording From Thousands of Neurons.”)
The device makes it possible to monitor single-neuron activity at an unprecedented scale, potentially opening new avenues for research. Investigators at the Allen Institute for Brain Science in Seattle, for example, are using the probe to expand some of the institute’s large-scale research tools — the Allen Cell Types Database, which characterizes cells from the mouse visual cortex, and the Allen Brain Observatory, which catalogs neuronal activity in cells from the same region.
Most efforts to catalog cells classify them according to anatomical, molecular or electrophysiological markers. But deciphering the link between these different modalities — whether molecular and electrophysiologically defined categories overlap, for example — has been challenging. Allen scientists aim to resolve that disconnect by systematically gathering large amounts of different kinds of information on the same neurons. Neuropixels’ ability to record electrical activity from hundreds to thousands of cells adds a new dimension to those data.
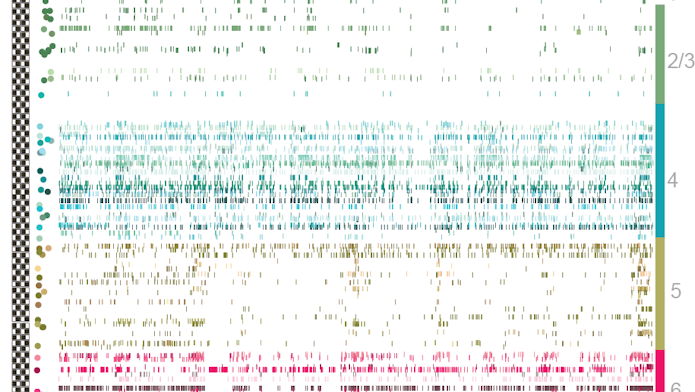
Dan Denman, a neuroscientist at the Allen Institute, and his collaborators are using extracellular field potentials recorded with Neuropixels from multiple sites around individual neurons to help catalog cells. Researchers look at how the shape of the action potentials changes across the cell, which in turn gives some clues to the cell’s morphology. They classify different features of the cells using machine learning and other techniques and then cluster each cell into a putative morphological class.
Early results suggest that cells in different regions can be distinguished by the shape of their action potentials, though Denman cautions that the results are still preliminary. “We can get enough information from the probe to tell us where the cell was, at least for cortical and subcortical structures,” Denman says. “This is all very early — they are not cell classes — but we think with more data it will be a powerful application.”
Researchers at the Allen Institute are also using Neuropixels to expand the Allen Brain Observatory, a database of neural activity recorded as mice passively view different stimuli. The Observatory has largely been focused on optical imaging, but Denman and his collaborators plan to replicate calcium imaging experiments using Neuropixels. That will make it possible to compare calcium imaging, an indirect measure of electrical activity, with electrophysiology, which has higher temporal resolution and a better signal-to-noise ratio.
“The relationship between calcium transients and action potentials is not 1-to-1 and can be complicated by cell type and conditions,” Denman says. “Having a more direct measure of action potentials is powerful.” Using the two technologies together will give researchers a better sense of how individual neurons are behaving with respect to the broader population.
Denman is also using Neuropixels to examine how individual cells’ activity can vary as the animal views the same stimuli over and over again, known as trial-to-trial variability. Preliminary research suggests that when an individual cell in the lateral geniculate nucleus (LGN) or primary visual cortex (V1) fires earlier than average in a given trial, the population within that area fires earlier than average as well. Variability across the two areas is only weakly correlated, however, suggesting that V1 does not solely inherit its variability from the LGN.
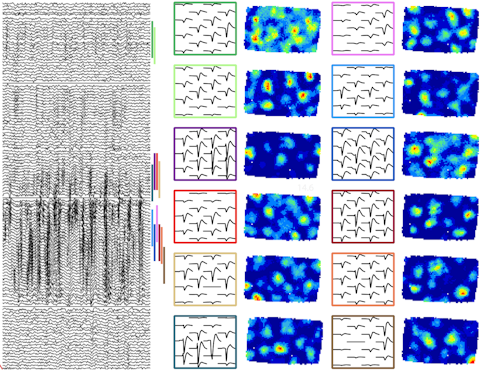
Marius Bauza, a postdoctoral scientist in John O’Keefe’s lab at University College London, is using Neuropixels to study grid cells. These cells are best known for their hexagonal receptive fields, which tile an animal’s environment and are thought to provide a sort of ruler for mapping space.
Bauza and his collaborators showed in 2015 that grid cell patterns can vary with the surroundings, potentially warping the spatial ruler. Put an animal in a trapezoid-shaped box, and cells that tile the slanted wall will lose their regular hexagonal receptive fields. That change raises an intriguing question, Bauza says. Can a combination of grid cells, some with irregular receptive fields, still properly map the environment?
Answering this question requires recording from ensembles of cells over time. Enter Neuropixels. Researchers are using the probe to study how receptive fields change when they move one wall in an animal’s environment. “To understand how the transformation takes place, you need to record from tens of cells, because position is encoded by the ensemble, not separate grid cells,” Bauza says.
By analyzing many cells simultaneously, the researchers developed a model for how receptive fields change with distance from the wall. “We found decoding could still be mediated by grid cell ensembles,” Bauza says. ‘Even distorted grid cells could provide a metric system.” The researchers are now looking at how interactions between cells in different brain areas give rise to the perception of space.


