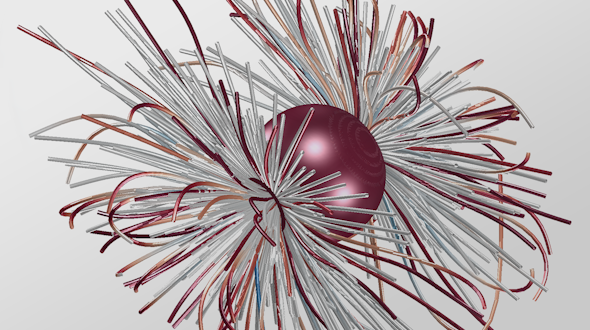
The Biophysical Modeling group focuses on the modeling and simulation of complex systems that arise in biology and soft condensed matter physics. Areas of interest include the dynamics of complex and active materials, as well as aspects of collective behavior and self-organization in both natural systems (e.g., inside the cell) and synthetic ones.
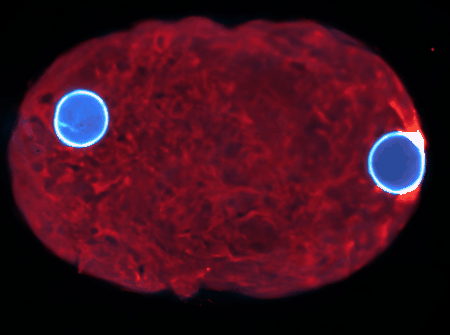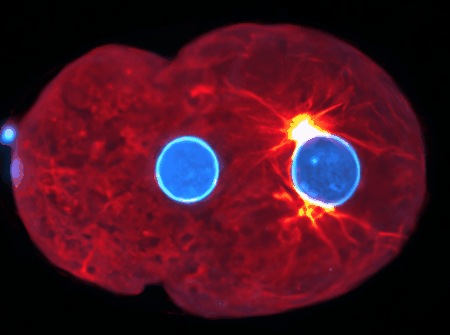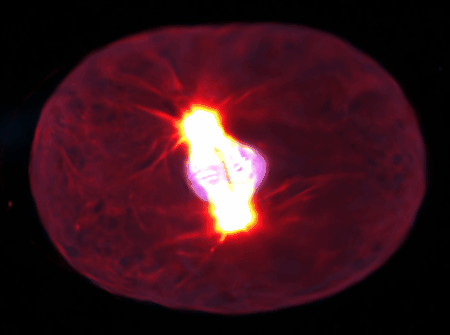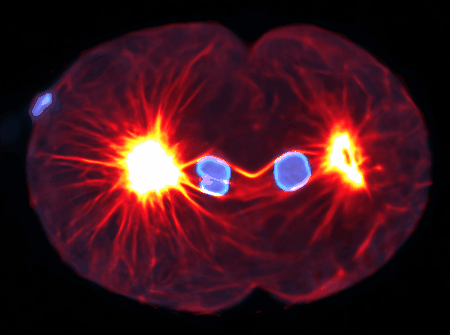
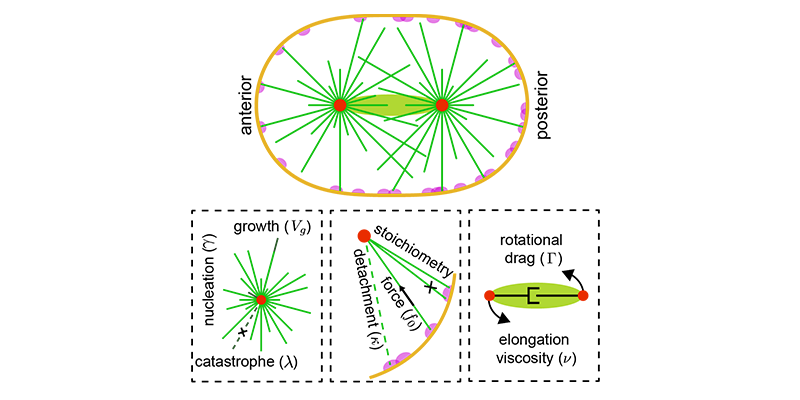
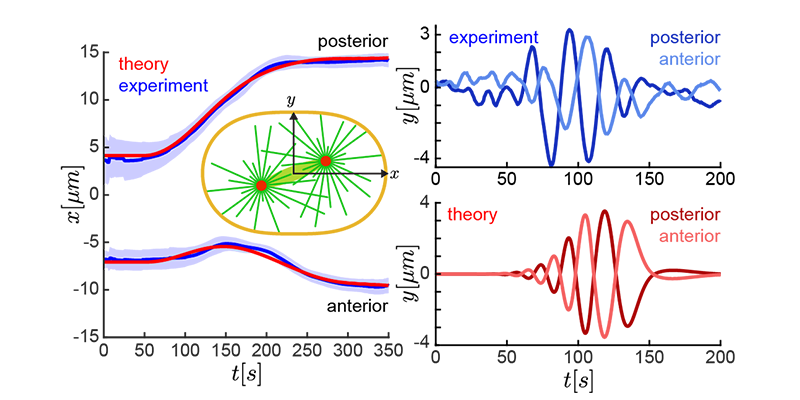
Examples are the organization and dynamics of the nucleus, the structure and assembly of spindles, the positioning and transport of cellular organelles, and fluid-structure problems in biology. To address these, often in close collaboration with experimental collaborators, we build numerical and theoretical models from the ground up, revealing how the known mechanics of individual components give rise to collective behavior. Many such phenomena occur only within dense, highly interacting systems, inaccessible to standard techniques. To probe such regimes requires the development of fast and scalable algorithms for many-component systems, and of coarse-grained models that can be analyzed and simulated.
Biophysical Modeling Lab